Dovetail® FFPE Assay
Unlock Genomic Variants in Your FFPE Samples
Dovetail linked-read technology enables high-sensitivity, comprehensive detection of structural variants (SVs) from FFPE samples.
FFPE samples are the most abundant resource in cancer research, yet nucleic acid degradation makes them challenging for most genomic assays—especially when detecting structural variants. Dovetail FFPE Technology overcomes this challenge.
Dovetail Genomics offers an FFPE-compatible workflow for generating linked-read libraries. Dovetail linked-reads enable high sensitivity detection of SVs, including coding and non-coding genomic rearrangements, while maintaining standard short-read performance for other variant (SNV, INDEL, CNV) detection.
The Dovetail® Linked-Read Advantage
Compatible with FFPE Samples – No high-molecular-weight DNA required.
Consistent performance – Validated across a variety of tumor types and varying DNA quality.
Amplifies true-variant signal – Increases SV detection sensitivity and minimizes false positives.
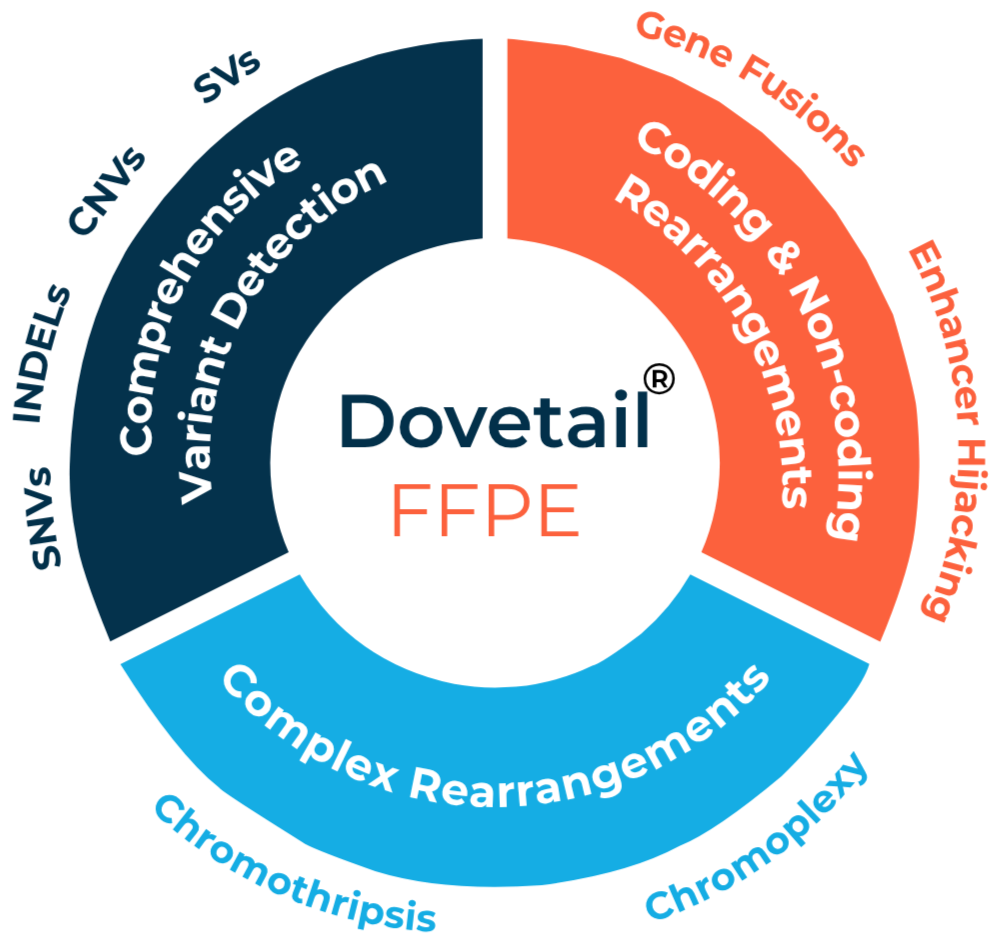
Comprehensive rearrangement profiling – Detects coding (gene fusions) and non-coding (enhancer hijacking) events.
Streamlines variant detection – Detects SV, SNVs, INDELs, and CNVs in a single assay.
Fits existing NGS infrastructure – Compatible with standard short-read platforms.
Dovetail FFPE technology is available as a kit or service- whichever works best for you.
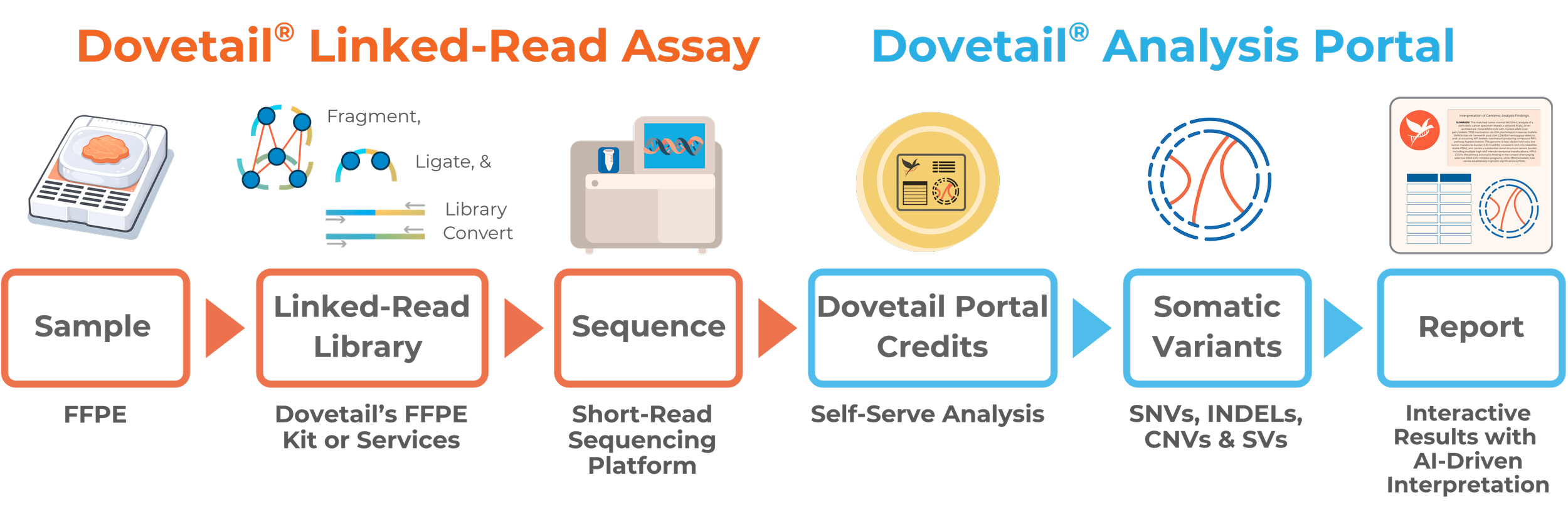
Dovetail addresses both library preparation and downstream data analysis in a single streamlined workflow. Prepare your library, sequence, and use the The Dovetail® Analysis Portal to upload your FASTQ files and submit the analysis in just a few clicks—no bioinformatics setup required. Receive results in a conveniently packaged downloadable archive, access a summary report with AI-driven interpretation of key findings, and explore your data with interactive tools.
A Complete Solution - Sample to Insight
Known Challenges With FFPE Samples
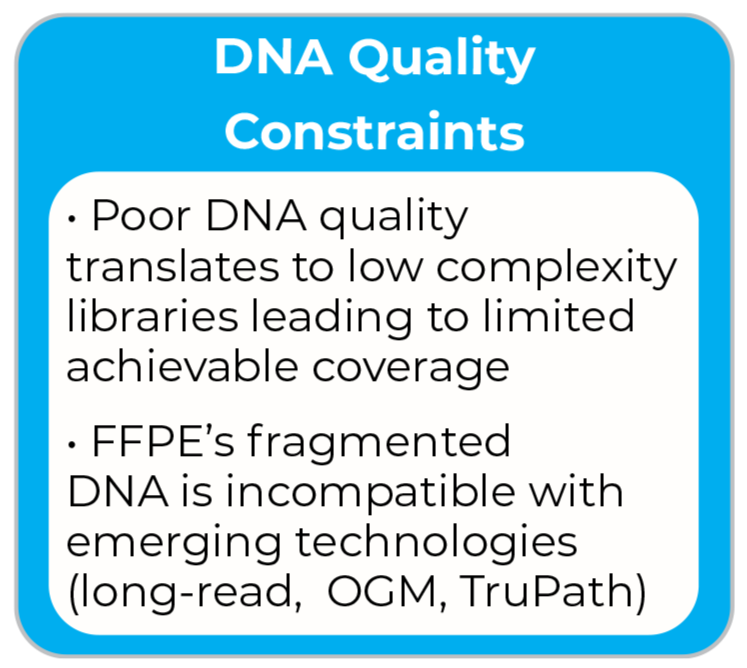



DNA Quality Constraints
· Poor DNA quality translates to low complexity libraries leading to limited achievable coverage
· FFPE’s fragmented DNA is incompatible with emerging technologies (long-reads, OGM, TruPath)
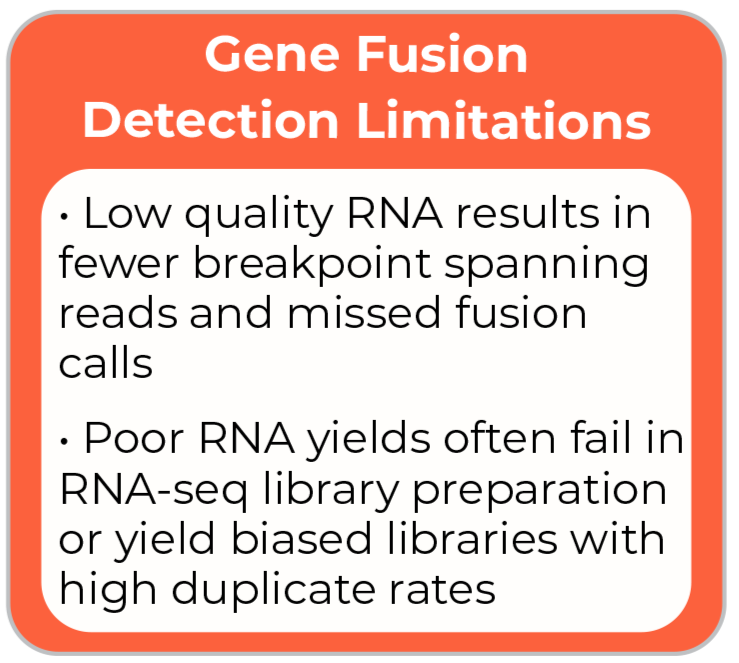
Gene Fusion Detection Limitations
· Low quality RNA results in fewer breakpoint spanning reads and missed fusion calls
· Poor RNA yields often fail in RNA-seq library preparation or yield biased, high duplicate libraries
High False Positive SV Calls
· Artifactual chimeric reads introduced during library prep are high in FFPE samples and inflate false positive SV calls
· Coverage drop-outs and spikes, common to FFPE NGS data, confuse SV callers
The Dovetail FFPE Solution
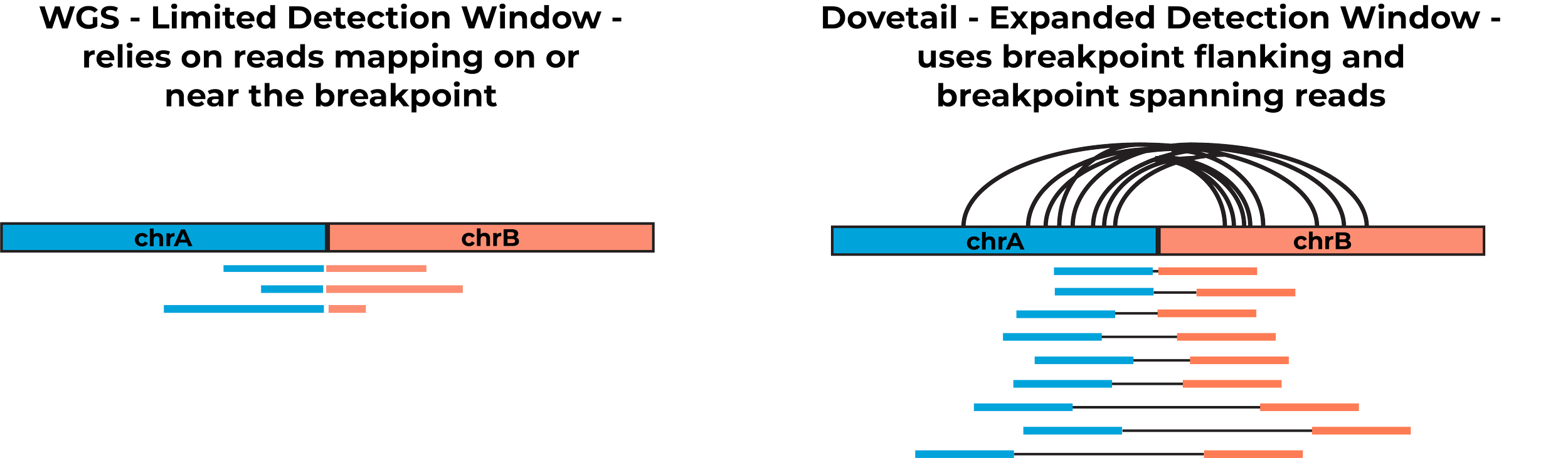
In standard whole genome sequencing, only reads that align to structural variant (SV) breakpoints support variant calls. Insufficient breakpoint-spanning reads —an issue intensified by limited achievable coverage in FFPE samples—lead to decreased sensitivity and increased false positives.
Dovetail technology overcomes these limitations by generating libraries that leverage both breakpoint-spanning and breakpoint-flanking reads, expanding the detection window to megabase-scale regions and significantly boosting sensitivity. This enables highly sensitive SV calling while maintaining WGS-equivalent performance for small variants, including SNVs, INDELs, and CNVs.
➜ Robust Performance
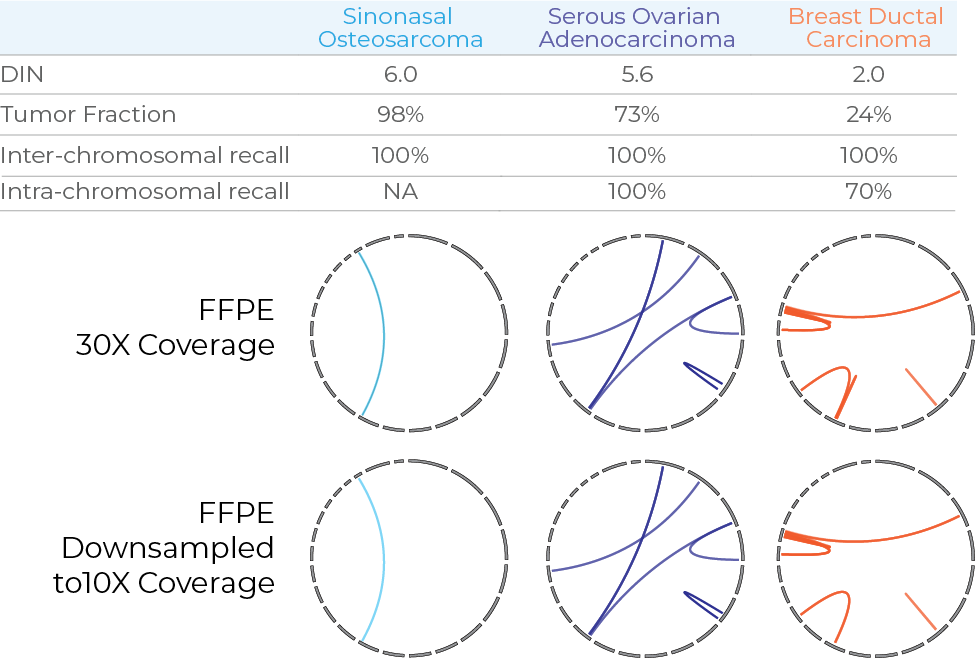
The Dovetail FFPE assay was used to prepare libraries from FFPE samples of varying quality, tumor types and fractions. Libraries were sequenced to 30X genome coverage and used to call both intra- and inter-chromosomal structural variants. Dovetail FFPE demonstrated robust performance across all sample types. Performance remained robust when the data was downsampled to 10X coverage.
➜High Concordance
The tumor FFPE samples profiled above were also available as fresh frozen tissue. Libraries were prepared from the fresh frozen samples and sequenced to equivalent coverage. Structural variants were called and variant allele fraction (VAF) was assigned to inter-chromosomal calls using the Dovetail Analysis Portal. SV calls between fresh frozen and FFPE samples showed high concordance in both breakpoint precision and VAF estimation.
PN: 20109V
Dovetail® FFPE Kit Full Solution, 8 Samples, 8 Libraries
Includes:
Dovetail® FFPE Kit
Dovetail® Library Module
Dovetail® Primer Module Set 1
Run the Dovetail® FFPE Kit in your lab
-
-
User Guides
-
Dovetail® FFPE Kit, 8 reactions
Includes:
Dovetail® Proximity Ligation Core
Dovetail® FFPE Module
Dovetail® FFPE Conditioning Module
Dovetail® Primer Module Set 1
Dovetail® Library Module
Gain access to our dedicated customer support team to guide your in-house workflows.
Leverage our specialist service team
Explore our optimized services to uncover critical genomic variants.
-
The Dovetail® FFPE Service delivers highly sensitive structural variant detection while preserving insights into all other variant types from your FFPE samples.
You provide:
11 × 5 µm scrolls (>20% tumor fraction)We provide:
Library preparation for you to carry out sequencing in-house or a sequenced library.A dedicated project manager will oversee your project and ensure smooth delivery.
Analysis is then available as a self-service option via the Dovetail Analysis Portal.
Turnaround time:
4 weeks -
Flexible deliverables tailored to your workflow:
Prefer to sequence in-house?
We can deliver sequencing-ready libraries to your lab.Lack sequencing capacity?
We can take your samples through sequencing and deliver raw data (FASTQ files).Need downstream insights?
Analysis is available as a self-service option via the Dovetail Analysis Portal.
Streamline Your Data Analysis
Analysis is available as a self-service option via the Dovetail Analysis Portal

The Dovetail Analysis Portal (DAP) offers customers instant access to streamlined, cloud-based analysis of NGS FASTQ files generated from Dovetail FFPE libraries. Accepting standard paired-end FASTQ inputs, it delivers packaged, easily downloadable results. Data analysis requires Dovetail Portal Credits.
Detect All Variant Classes
Identify SVs with high sensitivity
Alongside other variants: SNVs, INDELs, and CNVs
Access Results Easily
Insightful summary report
Data visualization
Downloadable outputs in standard formats for further exploration
Exploit Proprietary Analytics
Single base breakpoint refinement
Variant allele fraction (VAF) estimation including SVs
ecDNA vs. HSR predictor
Al-based interpretation of variant calls
Need our FFPE solution at scale?
Working with hundreds of samples?
Get in touch to explore partnership options.
We look forward to hearing from you!